Control of gene expression in prokaryotes and eukaryotes
By: David Balogh
Curriculum expectations:
D2.1 Use appropriate terminology related to molecular genetics
D3.2 Compare the structures and functions of RNA and DNA, and explain their roles in the process of protein synthesis
D3.3 Explain the steps involved in the process of protein synthesis and how genetic expression is controlled in prokaryotes and eukaryotes by regulatory proteins (e.g., the role of operons in prokaryotic cells; the mechanism of gene expression in eukaryotic cells)
D3.2 Compare the structures and functions of RNA and DNA, and explain their roles in the process of protein synthesis
D3.3 Explain the steps involved in the process of protein synthesis and how genetic expression is controlled in prokaryotes and eukaryotes by regulatory proteins (e.g., the role of operons in prokaryotic cells; the mechanism of gene expression in eukaryotic cells)
Learning Goals
Students will:
- Explain how genes are expressed through transcription and translation
- Explain the importance of controlling gene expression for both prokaryotic and eukaryotic organisms
- Explain the role of an operon in prokaryotic control of gene expression
- Use a simulated strand of DNA to demonstrate histone acetylation and methylation
- Use appropriate terminology to explain how histone acetylation and methylation makes DNA accessible and inaccessible (respectively) for transcription
- Create a model of acetylated and methylated strands of DNA
- Explain why histone acetylation and methylation is important in terms of gene expression
- Define the connection between telomeres and the aging process
control of gene expression
WHAT IS GENE EXPRESSION?
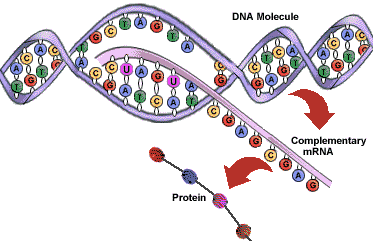
Gene expression is the process by which the information encoded into a gene is converted into a gene product, such as a protein or functional RNA. There are several steps in the process of gene expression, including transcription, RNA splicing, translation, and post-translational modifications. Gene expression can be controlled (and is) at any and all of these levels.
Gene expression is the process by which the information encoded into a gene is converted into a gene product, such as a protein or functional RNA. There are several steps in the process of gene expression, including transcription, RNA splicing, translation, and post-translational modifications. Gene expression can be controlled (and is) at any and all of these levels.
WHY IS CONTROLLING GENE EXPRESSION IMPORTANT?
Controlling how many and which types of genes are expressed gives cells the ability to determine their own structure and function. For example, in some multicellular organisms, such as many eukaryotes, a muscle cell may have vastly different requirements in terms of the proteins that it needs to function than a neuron. Similarly, single celled organisms such as many prokaryotes will need to express their genes in specific proportions to ensure their own survival and functionality.
Controlling how many and which types of genes are expressed gives cells the ability to determine their own structure and function. For example, in some multicellular organisms, such as many eukaryotes, a muscle cell may have vastly different requirements in terms of the proteins that it needs to function than a neuron. Similarly, single celled organisms such as many prokaryotes will need to express their genes in specific proportions to ensure their own survival and functionality.
Prokaryotes
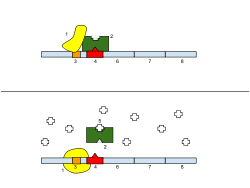
Prokaryotes tend to control their gene expression at the transcriptional level, often through the use of operons. An operon is a group of genes that functions together and is regulated together. A classic example of this is the lac operon, which was first discovered in e. coli. The lac operon in e. coli is responsible for regulating the expression of enzymes that break down lactose. Logically, the level of expression of these enzymes hinges on the amount of lactose present in the cell.
Under normal conditions, there is a lac repressor protein bound to the operator site of the gene sequence, which is located between the promoter region and the first gene of the operon. This repressor prevents RNA polymerase from transcribing the lactose gene, by blocking its path.
When lactose is introduced into the environment (and there is a sudden need for the production of enzymes to break it down), the lactose binds to the repressor, changing its conformation, and thus preventing it from binding to the DNA and blocking the path of RNA polymerase. This allows transcription to occur.
Under normal conditions, there is a lac repressor protein bound to the operator site of the gene sequence, which is located between the promoter region and the first gene of the operon. This repressor prevents RNA polymerase from transcribing the lactose gene, by blocking its path.
When lactose is introduced into the environment (and there is a sudden need for the production of enzymes to break it down), the lactose binds to the repressor, changing its conformation, and thus preventing it from binding to the DNA and blocking the path of RNA polymerase. This allows transcription to occur.
eukaryotes
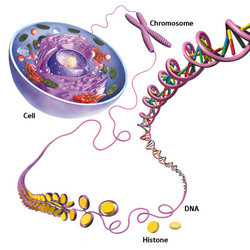
HISTONE MODIFICATIONS
In eukaryotes, gene expression can be controlled at a number of levels. At the transcriptional level, it can be regulated through histone modifications, which affect accessibility of the DNA by transcriptional machinery. Histones are proteins which form the core of each nucleosome, which is the main structural component of chromatin.
Histones can be chemically modified in a number of ways, which drastically affects how tightly the DNA is able to wind around the histones, and thus how tightly packed the chromatin is. Tightly packed chromatin, or heterochromatin, is inaccessible to transcriptional machinery. Loosely wound chromatin, or euchromatin, is accessible to transcriptional machinery.
Two examples of histone modifications are methylation and acetylation. Histone methylation involves the addition of one to three methyl groups to a histone, which can either repress or activate gene expression depending upon which histone is being modified, and how. Histone acetylation involves the addition of an acetyl group to a histone, which loosens the chromatin structure and makes the DNA accessible, thus increasing gene expression.
In eukaryotes, gene expression can be controlled at a number of levels. At the transcriptional level, it can be regulated through histone modifications, which affect accessibility of the DNA by transcriptional machinery. Histones are proteins which form the core of each nucleosome, which is the main structural component of chromatin.
Histones can be chemically modified in a number of ways, which drastically affects how tightly the DNA is able to wind around the histones, and thus how tightly packed the chromatin is. Tightly packed chromatin, or heterochromatin, is inaccessible to transcriptional machinery. Loosely wound chromatin, or euchromatin, is accessible to transcriptional machinery.
Two examples of histone modifications are methylation and acetylation. Histone methylation involves the addition of one to three methyl groups to a histone, which can either repress or activate gene expression depending upon which histone is being modified, and how. Histone acetylation involves the addition of an acetyl group to a histone, which loosens the chromatin structure and makes the DNA accessible, thus increasing gene expression.
Telomeres and aging
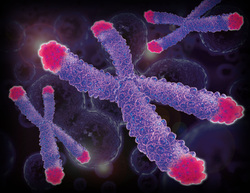
A telomere is a region of repetitive nucleotide sequences at each end of a chromatid, which contains the replicated DNA of a particular chromosome. Telomeres protect against deterioration of the chromosomes. Because the process of DNA replication is fundamentally reductive (some of the length of the chromosome is lost due to the nature of the replicative process), telomeres will inevitably shorten over the course of many replications. However, without telomeres, it would be the important "coding" regions of DNA that was being lost with each replication, which would obviously lead to a complete breakdown of the system.
Aging is thought to be a result of this limiting factor: eventually, the telomeres will shorten enough as to disappear completely, setting a limit on the number of times a chromosome can replicate, and thus the lifespan of an organism.
Aging is thought to be a result of this limiting factor: eventually, the telomeres will shorten enough as to disappear completely, setting a limit on the number of times a chromosome can replicate, and thus the lifespan of an organism.
Activities
| gene_expression_activity_handout.pdf | |
| File Size: | 5866 kb |
| File Type: | |
Spongelab games
ASSESSMENT
GLossary
Transcription: process by which DNA is copied into mRNA
Translation: process by which proteins are synthesized from mRNA
Eukaryote: organism whose cells contain complex, membrane-bound structures
Prokaryote: organism whose cells lack a membrane-bound nucleus
Operon: functioning unit of genomic DNA containing a cluster of genes that are expressed and regulated in tandem
Repressor: DNA-binding protein that regulates gene expression by blocking RNA polymerase from transcribing a gene
Histone: core structural component of a nucleosome, about which DNA is wound
Euchromatin: loosely packed, transcriptionally active DNA
Heterochromatin: tightly packed, transcriptionally inactive DNA
Methylation: introduction of a methyl functional group into a chemical compound
Acetylation: introduction of an acetyl functional group into a chemical compound
Telomere: region of repetitive nucleotide sequence at each end of a chromatid
Translation: process by which proteins are synthesized from mRNA
Eukaryote: organism whose cells contain complex, membrane-bound structures
Prokaryote: organism whose cells lack a membrane-bound nucleus
Operon: functioning unit of genomic DNA containing a cluster of genes that are expressed and regulated in tandem
Repressor: DNA-binding protein that regulates gene expression by blocking RNA polymerase from transcribing a gene
Histone: core structural component of a nucleosome, about which DNA is wound
Euchromatin: loosely packed, transcriptionally active DNA
Heterochromatin: tightly packed, transcriptionally inactive DNA
Methylation: introduction of a methyl functional group into a chemical compound
Acetylation: introduction of an acetyl functional group into a chemical compound
Telomere: region of repetitive nucleotide sequence at each end of a chromatid